Batch TriDFusion is an open-source software that can run multiple instances of the TriDFusion (3DF) workflow. The number of instances that can run simultaneously is customizable and can be adjusted according to the available GPU memory size, for example, in AI workflows.
Key Features
- Speed at scale: Processing large cohorts is significantly accelerated.
- Hardware-aware: Multiple systems can be used. Each system runs as many instances as its GPU memory allows (small, medium, or large GPUs all contribute).
- Consistent outputs: All results are consolidated into a unified, analysis-ready table.
- Simple operations: Define a protocol where series descriptions are used to sort the data, passing only the required series to the workflow, while controlling parallelism and monitoring progress.
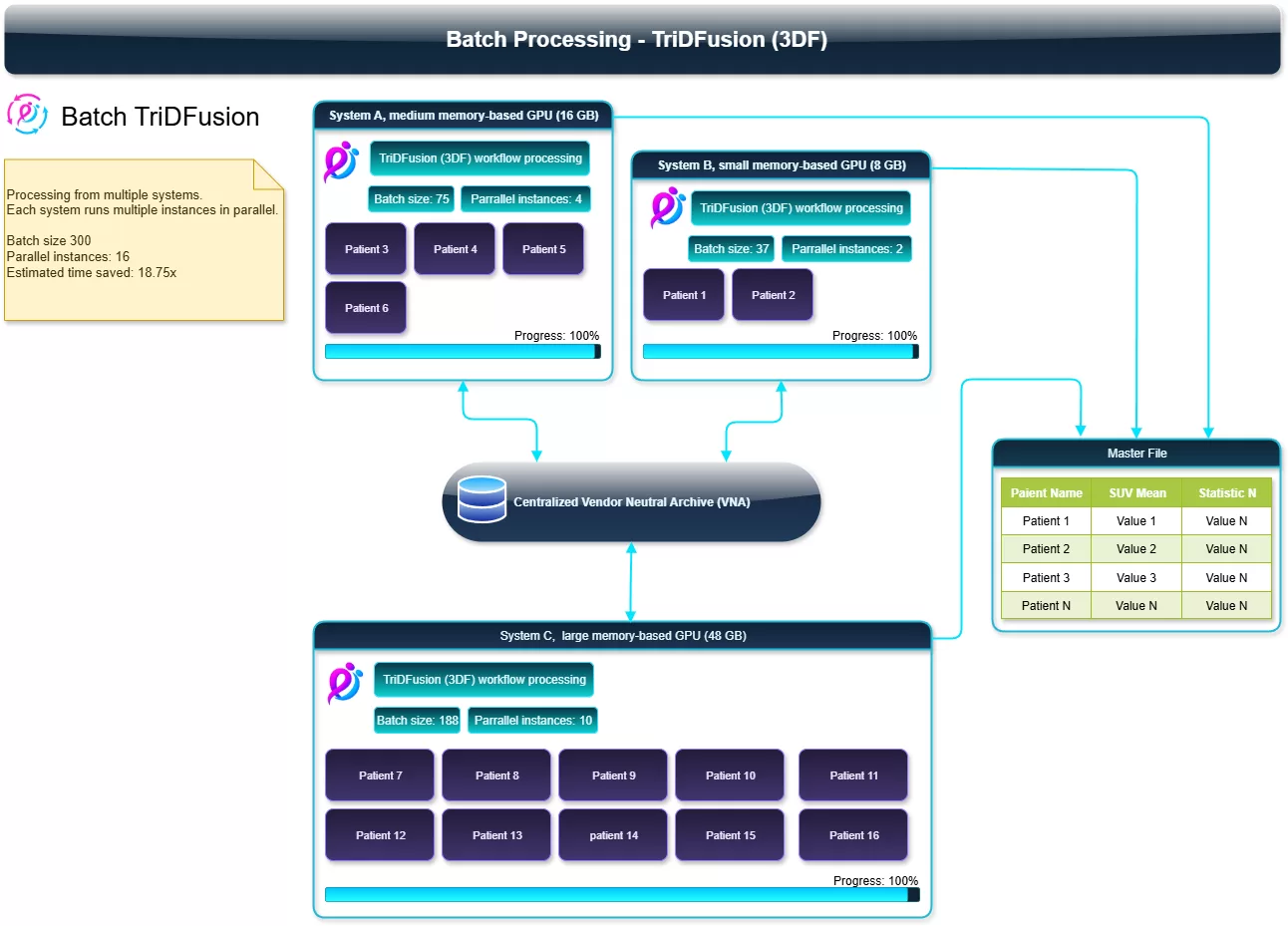
Example run (from the diagram)
- 3 systems participating (48 GB, 16 GB, 8 GB GPUs)
- 16 parallel instances total processing 300 patients
- Estimated time saved: ~18.75× vs. serial processing
- Output: master table (Patient Name, SUV Mean, additional statistics) in one place
Where it fits
- Large research cohorts where multiple workstations can be used to accelerate research outcomes.
Accessibility alt text for the diagram
Diagram of ‘TriDFusion (3DF) Batch Processing ’ showing three GPU systems (48 GB, 16 GB, 8 GB) each running several parallel instances. Arrows write results to a central VNA and into a master table (columns like Patient Name and SUV Mean). Callout notes: batch size 300, 16 parallel instances, ~18.75× time saved.
 Daniel Lafontaine
Daniel Lafontaine